This example shows how to plot pole figures using the built-in neml2 postprocessing regimes. Specifically, this example load a crystal plasticity model, uses that model to generate reorientation data, and then plots two types of pole figures: discrete points and a contour plot from a reconstructed Orientation Distribution Function (ODF). In the process then this also demonstrates how to reconstruct a continuous ODF from discrete data.
Importing required libraries and setting up
This example will run on the GPU with CUDA if available. nchunk is the chunk size for the pyzag integration, ncrystal is the number of random initial crystal orientations to use.
This example simulates rolling in an FCC material. rate sets the strain rate, total_rolling_strain the reduction strain, and ntime the number of time steps to take.
import matplotlib.pyplot as plt
import torch
import neml2
import neml2.tensors
from pyzag import nonlinear, chunktime
import neml2.postprocessing
torch.set_default_dtype(torch.double)
if torch.cuda.is_available():
dev = "cuda:0"
else:
dev = "cpu"
device = torch.device(dev)
nchunk = 20
ncrystal = 500
rate = 0.0001
total_rolling_strain = 0.5
ntime = 2500
end_time = total_rolling_strain / rate
static Rot rand(const TensorOptions &options=default_tensor_options())
Definition TransientDriver.h:32
deformation_rate = torch.zeros((ntime, ncrystal, 6), device = device)
deformation_rate[:, :, 1] = rate
deformation_rate[:, :, 2] = -rate
times = torch.linspace(0, end_time, ntime, device = device).unsqueeze(-1).unsqueeze(-1).expand((ntime, ncrystal, 1))
vorticity = torch.zeros((ntime, ncrystal, 3), device = device)
pyzag driver
For more details on the pyzag driver used here see the deterministic and stochastic inference examples. This helper object just lets us integrate the crystal model using pyzag
class Solve(torch.nn.Module):
"""Just integrate the model through some strain history
Args:
discrete_equations: the pyzag wrapped model
nchunk (int): number of vectorized time steps
rtol (float): relative tolerance to use for Newton's method during time integration
atol (float): absolute tolerance to use for Newton's method during time integration
"""
def __init__(self, discrete_equations, nchunk=1, rtol=1.0e-6, atol=1.0e-8):
super().__init__()
self.discrete_equations = discrete_equations
self.nchunk = nchunk
self.rtol = rtol
self.atol = atol
def forward(self, time, deformation_rate, vorticity, cache=False, initial_orientations=None):
"""Integrate through some time/temperature/strain history and return stress
Args:
time (torch.tensor): batched times
deformation_rate (torch.tensor): batched deformation rates
vorticity (torch.tensor): batched vocticities
Keyword Args:
cache (bool): if true, cache the solution and use it as a predictor for the next call.
This heuristic can speed things up during inference where the model is called repeatedly with similar parameter values.
"""
solver = nonlinear.RecursiveNonlinearEquationSolver(
self.discrete_equations,
step_generator=nonlinear.StepGenerator(self.nchunk),
predictor=nonlinear.PreviousStepsPredictor(),
nonlinear_solver=chunktime.ChunkNewtonRaphson(rtol=self.rtol, atol=self.atol),
)
forces = {
"deformation_rate":
neml2.SR2(deformation_rate, 0),
"initial_orientation":
neml2.Rot(initial_orientations, 0),
}
forces = [forces[key] for key in self.discrete_equations.fvars]
forces = (
.assemble()
.tensors[0]
)
state0 = [
neml2.Tensor()] * len(self.discrete_equations.svars)
if initial_orientations is not None:
i = self.discrete_equations.svars.index("orientation")
state0 = (
.assemble()
.tensors[0]
)
result = nonlinear.solve_adjoint(solver, state0, len(forces), forces)
if cache:
self.cached_solution = result.detach().clone()
return result
Rotation stored as modified Rodrigues parameters.
Definition Rot.h:49
The symmetric second order tensor.
Definition SR2.h:46
Scalar.
Definition Scalar.h:38
A skew-symmetric second order tensor, represented as an axial vector.
Definition WR2.h:43
Sparse representation of a vector consisting of a list of tensors and their layout.
Definition SparseVector.h:36
Simulate rolling deformation
Simulate the rolling deformation and extract the final crystal orientations
nmodel = neml2.load_nonlinear_system("crystal.i", "eq_sys")
nmodel.to(device=device)
model = Solve(neml2.pyzag.NEML2PyzagModel(nmodel), nchunk=nchunk)
with torch.no_grad():
orientations = neml2.tensors.Rot(
model(times, deformation_rate, vorticity, initial_orientations=initial_orientations)[
-1, :, 6:9
]
)
Plot a discrete pole figure
Use the built-in routines to plot a discerete 111 pole figure of the final orientations.
neml2.postprocessing.pretty_plot_pole_figure_points(
orientations, torch.tensor([1, 1, 1.0], device=device), crystal_symmetry="432"
)

png
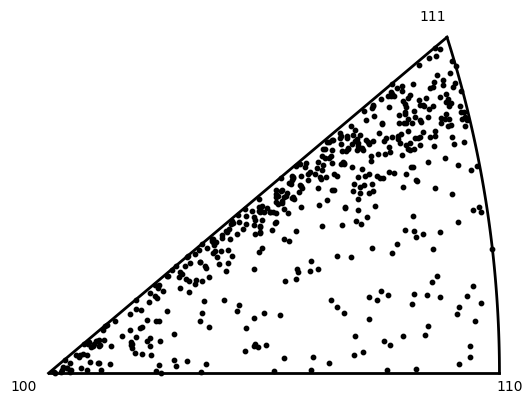
Plot a discrete inverse pole figure
Use the built-in routines to plot a discrete IPF in the rolling direction
neml2.postprocessing.pretty_plot_inverse_pole_figure(
orientations, torch.tensor([0.0, 1.0, 0.0], device=device), crystal_symmetry="432"
)
png
ODF reconstruction
Reconstruct the ODF from the discrete data. This example optimizes the kernel half-width with the build in routine (which uses a cross-validation approach). Print the final, optimal half width.
odf = neml2.postprocessing.odf.KDEODF(
orientations, neml2.postprocessing.odf.DeLaValleePoussinKernel(torch.tensor(0.1))
)
odf.optimize_kernel(verbose=True)
print(odf.kernel.h)
loss: -9.31230e-01: 100%|██████████| 50/50 [00:57<00:00, 1.15s/it]
Parameter containing:
tensor(0.1412, requires_grad=True)
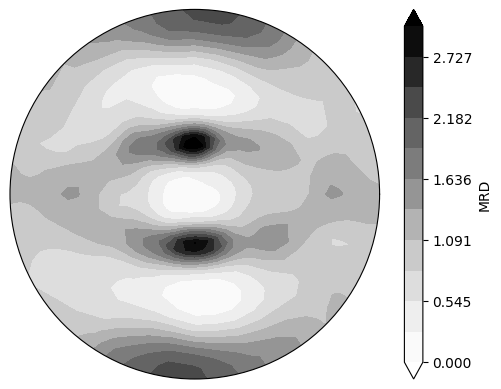
Plot a continuous polefigure
Use the reconstructed ODF to plot a continuous 111 polefigure
neml2.postprocessing.pretty_plot_pole_figure_odf(
odf,
torch.tensor([1, 1, 1.0], device=device),
crystal_symmetry="432",
limits=(0.0, 3.0),
ncontour=12,
)
png


