There are two common ways to formulate crystal plasticity kinematics:
- Integrate the elastic stretch and elastic rotation separately with two (coupled) rate equations.
- Integrate the plastic deformation gradient \(F_p\) directly from the plastic spatial velocity gradient \(l_p\).
Both formulations are based on the usual multiplicative decomposition of the deformation gradient into elastic and plastic parts \(F = F_e F_p\). This notebook calls the first approach the "separated" method and the second the "integrated" method (because the first splits the kinematics into two coupled equations and the second integrates a single equation for \(F_p\)).
It's easy to formulate a crystal model either way in NEML2 reusing most of detailed constitutive model objects like the hardening law, the slip law, and the crystal geometry. In fact the vast majority of the objects are shared between the two models with only a few specialized submodels required for each.
This example runs equivalent separated and integrated models through the same deformation history (rolling to 50% rolling strain) and plots a comparison. You can also use this notebook to look at differences in performance, though we have not done detailed performance optimization on either.
Importing required libraries and setting up
This example will run on the GPU with CUDA if available. nchunk is the chunk size for the pyzag integration, ncrystal is the number of random initial crystal orientations to use.
This example simulates rolling in an FCC material. rate sets the strain rate, total_rolling_strain the reduction strain, and ntime the number of time steps to take.
We need to calculate both the deformation rate/vorticity and the deformation gradient
import matplotlib.pyplot as plt
from matplotlib.lines import Line2D
import torch
import neml2
import neml2.tensors
from pyzag import nonlinear, chunktime
import neml2.postprocessing
torch.set_default_dtype(torch.double)
if torch.cuda.is_available():
dev = "cuda:0"
else:
dev = "cpu"
device = torch.device(dev)
nchunk = 10
ncrystal = 500
rate = 0.0001
total_rolling_strain = 0.5
ntime = 2500
end_time = total_rolling_strain / rate
initial_orientations = neml2.tensors.Rot.rand((ncrystal,), ()).
torch().to(device)
Definition TransientDriver.h:32
deformation_rate = torch.zeros((ntime, ncrystal, 6), device=device)
deformation_rate[:, :, 1] = rate
deformation_rate[:, :, 2] = -rate
times = (
torch.linspace(0, end_time, ntime, device=device)
.unsqueeze(-1)
.unsqueeze(-1)
.expand((ntime, ncrystal, 1))
)
vorticity = torch.zeros((ntime, ncrystal, 3), device=device)
full_spatial_velocity_gradient = (
neml2.tensors.R2(neml2.tensors.SR2(deformation_rate)).
torch()
+ neml2.tensors.R2(neml2.tensors.WR2(vorticity)).
torch()
)
F = torch.zeros_like(full_spatial_velocity_gradient)
F[0] = torch.eye(3, device=device)
dt = torch.diff(times, dim=0)
Finc = torch.linalg.matrix_exp(full_spatial_velocity_gradient[:-1] * dt.unsqueeze(-1))
for i in range(1, ntime):
F[i] = Finc[i - 1] @ F[i - 1]
pyzag driver for the separated model
For more details on the pyzag driver used here see the deterministic and stochastic inference examples. This helper object just lets us integrate the crystal model using pyzag
class SolveSeparate(torch.nn.Module):
"""Just integrate the model through some strain history
Args:
discrete_equations: the pyzag wrapped model
nchunk (int): number of vectorized time steps
rtol (float): relative tolerance to use for Newton's method during time integration
atol (float): absolute tolerance to use for Newton's method during time integration
"""
def __init__(self, discrete_equations, nchunk=1, rtol=1.0e-6, atol=1.0e-8):
super().__init__()
self.discrete_equations = discrete_equations
self.nchunk = nchunk
self.rtol = rtol
self.atol = atol
def forward(self, time, deformation_rate, vorticity, initial_orientations=None):
"""Integrate through some time/temperature/strain history and return stress
Args:
time (torch.tensor): batched times
deformation_rate (torch.tensor): batched deformation rates
vorticity (torch.tensor): batched vocticities
Keyword Args:
initial_orientation (torch.tensor): if provided, the initial orientations for each crystal
"""
solver = nonlinear.RecursiveNonlinearEquationSolver(
self.discrete_equations,
step_generator=nonlinear.StepGenerator(self.nchunk),
predictor=nonlinear.PreviousStepsPredictor(),
nonlinear_solver=chunktime.ChunkNewtonRaphson(rtol=self.rtol, atol=self.atol),
)
forces = {
"deformation_rate":
neml2.SR2(deformation_rate, 0),
"initial_orientation":
neml2.Rot(initial_orientations, 0),
}
forces = [forces[key] for key in self.discrete_equations.fvars]
forces = (
.assemble()
.tensors[0]
)
state0 = [
neml2.Tensor()] * len(self.discrete_equations.svars)
if initial_orientations is not None:
i = self.discrete_equations.svars.index("orientation")
state0 = (
.assemble()
.tensors[0]
)
result = nonlinear.solve_adjoint(solver, state0, len(forces), forces)
return result
Rotation stored as modified Rodrigues parameters.
Definition Rot.h:49
The symmetric second order tensor.
Definition SR2.h:46
Scalar.
Definition Scalar.h:38
A skew-symmetric second order tensor, represented as an axial vector.
Definition WR2.h:43
Sparse representation of a vector consisting of a list of tensors and their layout.
Definition SparseVector.h:36
Simulate rolling deformation for the seperated model
Simulate the rolling deformation and extract the final crystal orientations
nmodel_separate = neml2.load_nonlinear_system("crystal.i", "eq_sys")
nmodel_separate.to(device=device)
model_separate = SolveSeparate(neml2.pyzag.NEML2PyzagModel(nmodel_separate), nchunk=nchunk)
with torch.no_grad():
results_seperate = model_separate(
times, deformation_rate, vorticity, initial_orientations=initial_orientations
)
orientations_separate = neml2.tensors.Rot(results_seperate[-1, :, 6:9])
ODF reconstruction for the seperated model
Reconstruct the ODF from the discrete data. Uncommenting the two lines will optimize the kernel half-width with the built in routine (which uses a cross-validation approach). However, for consistency between the two methods, we'll use a fixed half width for both.
odf_separate = neml2.postprocessing.odf.KDEODF(
orientations_separate, neml2.postprocessing.odf.DeLaValleePoussinKernel(torch.tensor(0.2))
)
odf_separate.kernel.h = torch.tensor(0.23)
pyzag driver for the integrated model
Same thing as for the separated model, just now for the integrated form where we take the deformation gradient as input. Don't forget to set the initial plastic deformation gradient to the identity!
class SolveIntegrated(torch.nn.Module):
"""Just integrate the model through some strain history
Args:
discrete_equations: the pyzag wrapped model
nchunk (int): number of vectorized time steps
rtol (float): relative tolerance to use for Newton's method during time integration
atol (float): absolute tolerance to use for Newton's method during time integration
"""
def __init__(self, discrete_equations, nchunk=1, rtol=1.0e-6, atol=1.0e-8):
super().__init__()
self.discrete_equations = discrete_equations
self.nchunk = nchunk
self.rtol = rtol
self.atol = atol
def forward(self, time, deformation_gradient, initial_orientations=None):
"""Integrate through some time/temperature/strain history and return stress
Args:
time (torch.tensor): batched times
deformation_gradient (torch.tensor): batched deformation gradients
Keyword Args:
initial_orientation (torch.tensor): if provided, the initial orientations for each crystal
"""
solver = nonlinear.RecursiveNonlinearEquationSolver(
self.discrete_equations,
step_generator=nonlinear.StepGenerator(self.nchunk),
predictor=nonlinear.PreviousStepsPredictor(),
nonlinear_solver=chunktime.ChunkNewtonRaphson(rtol=self.rtol, atol=self.atol),
)
forces = {
}
forces = [forces[key] for key in self.discrete_equations.fvars]
forces = (
.assemble()
.tensors[0]
)
state0 = [
neml2.Tensor()] * len(self.discrete_equations.svars)
i = self.discrete_equations.svars.index("Fp")
state0[i] = neml2.tensors.Tensor(
torch.eye(3, device=time.device).unsqueeze(0).expand(time.shape[1:2] + (3, 3)), 1
)
state0 = (
.assemble()
.tensors[0]
)
result = nonlinear.solve_adjoint(solver, state0, len(forces), forces)
return result
Base class for second order tensor.
Definition R2.h:49
Simulate for the multiplicative model
Again, simulate rolling deformation
nmodel_integrated = neml2.load_nonlinear_system("crystal_integrated.i", "eq_sys")
nmodel_integrated.to(device=device)
model_integrated = SolveIntegrated(neml2.pyzag.NEML2PyzagModel(nmodel_integrated), nchunk=nchunk)
with torch.no_grad():
results_integrated = model_integrated(times, F, initial_orientations=initial_orientations)
end_results_integrated = results_integrated[-1]
Extract the orientations for the multiplicative model
This takes some doing. We first need to get \(F_p\) from the state, then calculate \(F_e = F F_p^{-1}\), then do a polar decomposition to get the rotation as a matrix, then convert to modified Rodrigues parameters. Finally compose with the original texture to get the final, deformed texture.
split_state = (
model_integrated.discrete_equations.slayout, [
neml2.Tensor(end_results_integrated, 1)]
)
.disassemble()
.tensors
)
iFp = model_integrated.discrete_equations.svars.index("Fp")
Fp = split_state[iFp].
torch().reshape(-1, 3, 3)
Flast = F[-1]
Fe = Flast @ torch.linalg.inv(Fp)
U, S, Vh = torch.linalg.svd(Fe)
Re = U @ Vh
Ue = Vh.transpose(-2, -1).conj() @ torch.diag_embed(S, dim1=-2, dim2=-1) @ Vh
Re_mrp = neml2.tensors.Rot.fill_matrix(neml2.tensors.R2(Re))
Q0 = neml2.tensors.Rot(initial_orientations)
Q = Re_mrp * Q0
Dense representation of a tensor assembled from a list of tensors and their layout.
Definition AssembledVector.h:36
Construct the ODF for the multiplicative model results
Basically the same as for the separated approach.
odf_integrated = neml2.postprocessing.odf.KDEODF(
Q, neml2.postprocessing.odf.DeLaValleePoussinKernel(torch.tensor(0.2))
)
odf_integrated.kernel.h = torch.tensor(0.23)
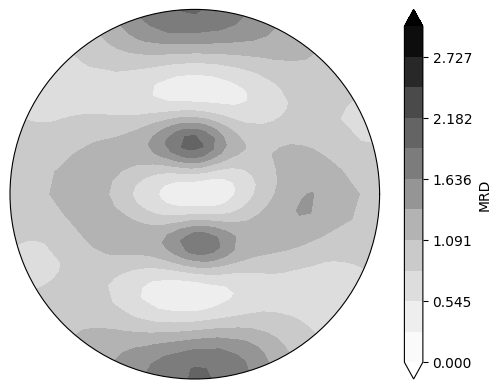
Plot the pole figures
Use the reconstructed ODFs to plot continuous 111 polefigure
neml2.postprocessing.pretty_plot_pole_figure_odf(
odf_separate,
torch.tensor([1, 1, 1.0], device=device),
crystal_symmetry="432",
limits=(0.0, 3.0),
ncontour=12,
)
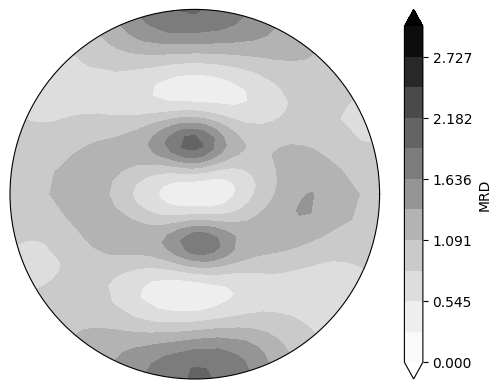
neml2.postprocessing.pretty_plot_pole_figure_odf(
odf_integrated,
torch.tensor([1, 1, 1.0], device=device),
crystal_symmetry="432",
limits=(0.0, 3.0),
ncontour=12,
)
png
png
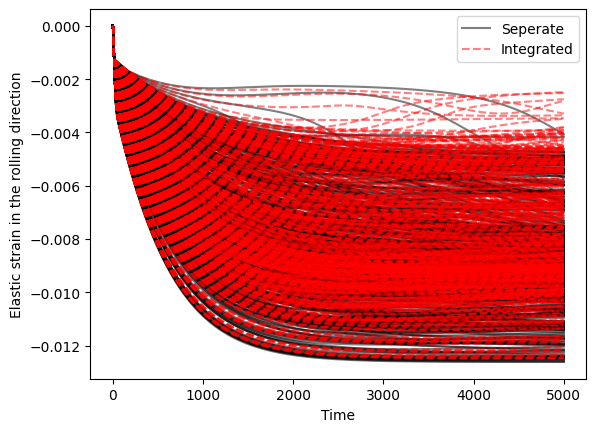
Plot some comparisons of the elastic strains
Extract the elastic strains from each model and plot:
- The history of each crystal over time
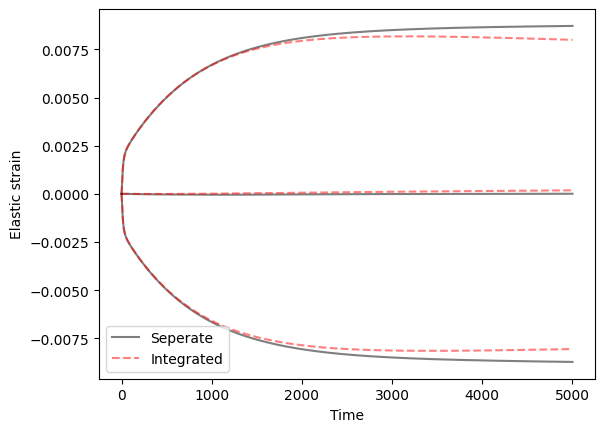
- The average over time
The averages are consistent but there are some differences in stress/elastic strain history between crystals when calculated this way.
state_history_split = (
model_separate.discrete_equations.slayout, [
neml2.Tensor(results_seperate, 2)]
)
.disassemble()
.tensors
)
state_history_mult = (
model_integrated.discrete_equations.slayout, [
neml2.Tensor(results_integrated, 2)]
)
.disassemble()
.tensors
)
iFp = model_integrated.discrete_equations.svars.index("Fp")
Fp_mult = state_history_mult[iFp].
torch().reshape(F.shape[:-2] + (3, 3))
Fe_mult = F @ torch.linalg.inv(Fp_mult)
U_mult, S_mult, Vh_mult = torch.linalg.svd(Fe_mult)
Re_mult = U_mult @ Vh_mult
Ue_mult = Vh_mult.transpose(-2, -1).conj() @ torch.diag_embed(S_mult, dim1=-2, dim2=-1) @ Vh_mult
E_mult = 0.5 * (Ue_mult.transpose(-2, -1) @ Ue_mult - torch.eye(3, device=device))
ie = model_separate.discrete_equations.svars.index("elastic_strain")
plt.plot(times[:, 0, 0].cpu(), state_history_split[ie].
torch()[:, :, 2].cpu(),
"k-", alpha=0.5)
plt.plot(times[:, 0, 0].cpu(), E_mult[:, :, 2, 2].cpu(), "r--", alpha=0.5)
plt.xlabel("Time")
plt.ylabel("Elastic strain in the rolling direction")
plt.legend(
[
Line2D([0], [0], color="k", alpha=0.5),
Line2D([0], [0], color="r", alpha=0.5, linestyle="--"),
],
["Seperate", "Integrated"],
loc="best",
)
<matplotlib.legend.Legend at 0x7253c41c6e70>
png
for i in range(3):
plt.plot(
times[:, 0, 0].cpu(),
torch.mean(state_history_split[ie].
torch()[:, :, i], dim=-1).cpu(),
"k-",
alpha=0.5,
)
plt.plot(times[:, 0, 0].cpu(), torch.mean(E_mult[:, :, i, i], dim=-1).cpu(), "r--", alpha=0.5)
plt.xlabel("Time")
plt.ylabel("Elastic strain")
plt.legend(
[
Line2D([0], [0], color="k", alpha=0.5),
Line2D([0], [0], color="r", alpha=0.5, linestyle="--"),
],
["Seperate", "Integrated"],
loc="best",
)
<matplotlib.legend.Legend at 0x7253c41c4cb0>
png



